 -
Volume 58,
Issue 2,
2008
-
Volume 58,
Issue 2,
2008
Volume 58, Issue 2, 2008
- Notification List
-
-
-
Notification that new names and new combinations have appeared in volume 57, part 11, of the IJSEM
More LessThis listing of names published in a previous issue of the IJSEM is provided as a service to bacteriology to assist in the recognition of new names and new combinations. This procedure was proposed by the Judicial Commission [Minute 11(ii), Int J Syst Bacteriol 41 (1991), p. 185]. The names given herein are listed according to the Rules of priority (i.e. page number and order of valid publication of names in the original articles). Taxonomic opinions included in this List (i.e. the creation of synonyms or the emendation of circumscriptions) cannot be considered as validly published nor, in any other way, approved by the International Committee on Systematics of Prokaryotes and its Judicial Commission.
-
-
- New Taxa
-
- Actinobacteria
-
-
Actinomycetospora chiangmaiensis gen. nov., sp. nov., a new member of the family Pseudonocardiaceae
More LessA novel actinomycete strain, YIM 0006T, was isolated from soil of a tropical rainforest in northern Thailand. The isolate displayed the following characteristics: aerial mycelium is absent, short spore chains are formed directly on the substrate mycelium, contains meso-diaminopimelic acid, arabinose and galactose (cell-wall chemotype IV), the diagnostic phospholipid is phosphatidylcholine, MK-9(H4) is the predominant menaquinone and the G+C content of the genomic DNA is 69.0 mol%. Phylogenetic analysis and phenotypic characteristics showed that strain YIM 0006T belongs to the family Pseudonocardiaceae but can be distinguished from representatives of all genera classified in the family. The novel genus and species Actinomycetospora chiangmaiensis gen. nov., sp. nov. are proposed, with strain YIM 0006T (=CCTCC AA 205017T =DSM 45062T) as the type strain of Actinomycetospora chiangmaiensis.
-
-
-
Microbacterium ginsengisoli sp. nov., a β-glucosidase-producing bacterium isolated from soil of a ginseng field
More LessStrain Gsoil 259T, a β-glucosidase-producing bacterium, was isolated from a soil sample from a ginseng field in the Republic of Korea and characterized in order to determine its taxonomic position. Cells were Gram-positive, heterotrophic, strictly aerobic, non-motile short rods. 16S rRNA gene sequence analysis revealed that strain Gsoil 259T belonged to the genus Microbacterium and was closely related to Microbacterium arborescens IFO 3750T (98.5 %) and Microbacterium imperiale IFO 12610T (97.9 %). However, it has low values for DNA–DNA relatedness with the above strains (20.7 and 17.5 %, respectively). Strain Gsoil 259T possessed chemotaxonomic markers that were consistent with classification in the genus Microbacterium, i.e. MK-11 and MK-12 were the major menaquinones and anteiso-C17 : 0, anteiso-C15 : 0 and iso-C16 : 0 were the predominant cellular fatty acids. The DNA G+C content was 69.4 mol%. The cell-wall sugar was rhamnose and the diamino acid in the cell-wall peptidoglycan was ornithine. On the basis of data from this polyphasic study, strain Gsoil 259T represents a novel species of the genus Microbacterium, for which the name Microbacterium ginsengisoli sp. nov. is proposed. The type strain is Gsoil 259T (=KCTC 19189T =DSM 18659T).
-
-
-
Nocardioides hankookensis sp. nov., isolated from soil
More LessA Gram-positive, non-motile, rod-shaped bacterial strain, DS-30T, was isolated from soil from Dokdo, Korea, and its taxonomic position was investigated by means of a polyphasic study. Strain DS-30T grew optimally at pH 6.0–7.0 and 25 °C in the presence of 0–0.5 % (w/v) NaCl. A phylogenetic analysis based on 16S rRNA gene sequences showed that strain DS-30T fell within the genus Nocardioides, forming a coherent cluster with the type strains of Nocardioides aquiterrae and Nocardioides pyridinolyticus, with which it exhibited 16S rRNA gene sequence similarity values of 98.1 and 98.3 %, respectively. Strain DS-30T exhibited 16S rRNA gene sequence similarity values below 96.8 % with respect to the type strains of the other Nocardioides species. The chemotaxonomic properties of strain DS-30T were consistent with those of the genus Nocardioides: the cell-wall peptidoglycan type was based on ll-diaminopimelic acid, the predominant menaquinone was MK-8(H4) and the major fatty acids were iso-C16 : 0, C17 : 1 ω8c and C18 : 1 ω9c. The DNA G+C content was 71.3 mol%. Strain DS-30T exhibited DNA–DNA relatedness levels of 19 and 22 % with respect to the type strains of N. aquiterrae and N. pyridinolyticus, respectively, and could be differentiated from these species with reference to a number of phenotypic characteristics. On the basis of the data obtained, strain DS-30T represents a novel species of the genus Nocardioides, for which the name Nocardioides hankookensis sp. nov. is proposed. The type strain is DS-30T (=KCTC 19246T =CCUG 54522T).
-
-
-
Kribbella hippodromi sp. nov., isolated from soil from a racecourse in South Africa
More LessA novel actinomycete, designated strain S1.4T, was isolated from a soil sample collected from Kenilworth Racecourse in the Western Cape, South Africa. The strain was able to grow in the presence of 5 % NaCl. It contained ll-diaminopimelic acid and glycine in the cell-wall peptidoglycan with glucose present in the whole-cell sugar profile. Strain S1.4T was shown to be a member of either the genus Kribbella or the genus Nocardioides based on a rapid molecular identification method by using single-enzyme restriction endonuclease digestion of the PCR-amplified 16S rRNA gene with MboI, VspI, SphI, SnaBI, SalI and AgeI. Analysis of the 16S rRNA gene sequence indicated that strain S1.4T belonged to the genus Kribbella. Phylogenetic analysis based on 16S rRNA gene sequence comparisons showed that strain S1.4T was related most closely to Kribbella solani DSA1T. Strain S1.4T was phenotypically distinct from K. solani DSA1T and was shown to be a separate genomic species based on DNA–DNA hybridization experiments (40.4±3.8 % DNA–DNA relatedness between the two). Strain S1.4T (=DSM 19227T =NRRL B-24553T) is thus presented as the type strain of a novel species, for which the name Kribbella hippodromi sp. nov. is proposed.
-
-
-
Mycobacterium setense sp. nov., a Mycobacterium fortuitum-group organism isolated from a patient with soft tissue infection and osteitis
More LessA Gram-positive, rod-shaped acid-fast bacterium was isolated from a patient with a post-traumatic chronic skin abscess associated with osteitis. Morphological analysis, 16S rRNA, hsp65, sodA and rpoB gene sequence analysis, cell-wall fatty acid and mycolic acid composition analyses and biochemical tests showed that the isolate, designated ABO-M06T, belonged to the genus Mycobacterium. Its phenotype was unique and genetic and phylogenetic findings suggest that strain ABO-M06T represents a novel species within the Mycobacterium fortuitum group. The name Mycobacterium setense sp. nov. is proposed for this novel species, with the type strain ABO-M06T (=CIP 109395T=DSM 45070T).
-
-
-
Brevibacterium marinum sp. nov., isolated from seawater
More LessA novel yellow-pigmented actinobacterium was isolated from seawater collected from Hwasun Beach in Jeju, Republic of Korea. A comparative analysis of the 16S rRNA gene sequence indicated that the organism, designated HFW-26T, was closely related to members of the genus Brevibacterium. As found for other species of the genus Brevibacterium, strain HFW-26T possessed meso-diaminopimelic acid as the diagnostic cell-wall diamino acid, contained MK-8(H2) as the major menaquinone, contained polar lipids that included diphosphatidylglycerol, phosphatidylglycerol, phosphatidylinositol and an unknown phospholipid, and had anteiso-C15 : 0 and anteiso-C17 : 0 as the predominant fatty acids. The G+C content of the DNA was 71.4 mol%. The phylogenetically closest relative was Brevibacterium picturae DSM 16132T (99.0 % 16S rRNA gene sequence similarity). However, DNA–DNA hybridization of strain HFW-26T showed 35.1–43.7 % relatedness with respect to B. picturae DSM 16132T. The novel isolate could be clearly distinguished from B. picturae DSM 16132T on the basis of some cultural, physiological and biochemical characteristics. A battery of phenotypic and genetic data obtained in this study suggest that strain HFW-26T represents a novel species of the genus Brevibacterium, for which the name Brevibacterium marinum sp. nov. is proposed. The type strain is HFW-26T (=JBRI 2001T=KCTC 19221T=DSM 18964T).
-
- Bacteroidetes
-
-
Gaetbulibacter marinus sp. nov., isolated from coastal seawater, and emended description of the genus Gaetbulibacter
More LessA Gram-negative, yellow-coloured, chemoheterotrophic, non-motile, strictly aerobic, rod-shaped bacterium, designated IMCC1914T, was isolated from coastal surface seawater of the Yellow Sea, Korea. The temperature, pH and NaCl ranges for growth were 3–37 °C, pH 8.0–11.0 and 0.5–4.0 %. The DNA G+C content of the strain was 38.1 mol% and the major cellular fatty acids were iso-C15 : 1 (32.1 %), iso-C15 : 0 (20.6 %) and iso-C17 : 0 3-OH (7.8 %). Phylogenetic analysis based on 16S rRNA gene sequences indicated that strain IMCC1914T was related most closely to Gaetbulibacter saemankumensis SMK-12T, with a sequence similarity of 96.2 %. On the basis of phylogenetic data and several distinct phenotypic characteristics, strain IMCC1914T (=KCCM 42380T =NBRC 102040T) could be assigned to the genus Gaetbulibacter as the type strain of a novel species, for which the name Gaetbulibacter marinus sp. nov. is proposed. In addition, an emended description of the genus Gaetbulibacter is presented.
-
-
-
Diversity of Capnocytophaga species in children and description of Capnocytophaga leadbetteri sp. nov. and Capnocytophaga genospecies AHN8471
More LessBacteria of the genus Capnocytophaga form part of the resident oral flora in children and adults. They are recognized as opportunistic pathogens of various extra-oral infections. The significance of individual species in periodontal and extra-oral diseases is unclear, due to the inability of conventional phenotypic tests to identify clinical isolates to species level. Aiming at a clear distinction between species, we undertook a phylogenetic study of a collection of 102 Capnocytophaga strains including 62 oral isolates from children and 40 reference strains from oral and extra-oral infections representing the five known, human, oral Capnocytophaga species. The phylogeny was estimated on the basis of multilocus enzyme electrophoresis (MLEE) of 12 intracellular, housekeeping enzymes and by partial 16S rRNA gene sequencing, and was compared to phenotypic characteristics. The clustering profiles in the MLEE and sequence-based dendrograms were concordant and allowed identification of isolates to species level, based on co-clustering with reference strains. The study confirmed Capnocytophaga ochracea and Capnocytophaga sputigena as separate species, and underlined the problems of distinguishing between them by conventional phenotypic tests. The presence of two distinct clusters of oral isolates from children indicated the existence of novel species, supported by analysis of near-full-length 16S rRNA gene sequences and by DNA–DNA hybridization results. One cluster of weakly saccharolytic isolates without the ability to ferment sucrose is proposed as Capnocytophaga leadbetteri sp. nov. (type strain AHN8855T=CCUG 51857T=NCTC 13375T). Another cluster not phenotypically distinguishable from C. ochracea and C. sputigena is designated Capnocytophaga genospecies AHN8471 (represented by strain AHN8471=CCUG 51856=NCTC 13374).
-
-
-
Parapedobacter soli sp. nov., isolated from soil of a ginseng field
More LessStrain DCY14T, a Gram-negative, non-spore-forming, rod-shaped, non-motile bacterium, was isolated from soil from a ginseng field in Korea and was characterized in order to determine its taxonomic position. 16S rRNA gene sequence analysis revealed that strain DCY14T belongs to the family Sphingobacteriaceae, the highest degree of sequence similarity being found with respect to Parapedobacter koreensis Jip14T (95.8 %). Chemotaxonomic data revealed that strain DCY14T possesses MK-7 as the major menaquinone. The major fatty acids present were anteiso-C13 : 0, iso-C15 : 0, iso-C17 : 0 3-OH and summed feature 4 (C16 : 1 ω7c/iso-C15 : 0 2-OH). The results of physiological and biochemical tests clearly demonstrated that strain DCY14T represents a distinct species. On the basis of these data, DCY14T represents a novel species of the genus Parapedobacter, for which the name Parapedobacter soli sp. nov. is proposed. The type strain is DCY14T (=KCTC 12984T =LMG 24069T).
-
-
-
Bacteroides propionicifaciens sp. nov., isolated from rice-straw residue in a methanogenic reactor treating waste from cattle farms
More LessTwo strictly anaerobic bacterial strains (SV434T and S562) were isolated from rice-straw residue in a methanogenic reactor treating waste from cattle farms in Japan. They had identical 16S rRNA gene sequences and showed almost the same phenotypic properties. The cells of both strains were Gram-negative, non-motile, non-spore-forming rods; extraordinarily long rods often occurred. Remarkable stimulation of growth occurred with the addition of haemin and cobalamin (vitamin B12) to the medium. The supplementary cobalamin and haemin could be replaced if autoclaved and clarified sludge fluid obtained from the reactor was added. Both strains utilized a range of growth substrates, including arabinose, fructose, galactose, glucose, mannose, cellobiose, maltose, glycogen, starch, dextrin, amygdalin, lactate and pyruvate. Both strains produced acetate and propionate with a small amount of succinate from these substrates in the presence of haemin and cobalamin. Both strains were slightly alkaliphilic, having a pH optimum at about 7.9. The temperature range for growth was 5–35 °C, the optimum being 30 °C. The NaCl concentration range for growth was 0–4 % (w/v). Catalase activity was not detected in cells cultivated without haemin, whereas cells cultivated with haemin usually had the enzyme activity. Oxidase and nitrate-reducing activities were not detected. Aesculin was hydrolysed, but gelatin was not hydrolysed. Both strains were sensitive to bile acids. The major cellular fatty acids of both strains were anteiso-C15 : 0 and iso-C15 : 0. Menaquinones MK-8(H0) and MK-9(H0) were the major respiratory quinones and the genomic DNA G+C contents were 46.2–47.5 mol%. A phylogenetic analysis based on 16S rRNA gene sequences placed both strains in the phylum Bacteroidetes. Bacteroides coprosuis (isolated from swine-manure storage pits) was the species most closely related to both strains (95.9 % 16S rRNA gene sequence similarity to the type strain). On the basis of the phylogenetic, physiological and chemotaxonomic analyses, strains SV434T and S562 represent a novel species of the genus Bacteroides, for which the name Bacteroides propionicifaciens sp. nov. is proposed. The type strain is SV434T (=JCM 14649T =DSM 19291T).
-
-
-
Salegentibacter salinarum sp. nov., isolated from a marine solar saltern
More LessA bacterial strain, ISL-4T, comprising Gram-negative, non-motile, rod-shaped cells, was isolated from a marine solar saltern of the Yellow Sea in Korea and was subjected to a polyphasic taxonomic investigation. Phenotypically, the strain resembled members of the genus Salegentibacter. It grew optimally at pH 7.0–8.0 and 30 °C and in the presence of 8 % (w/v) NaCl. It contained MK-6 as the predominant menaquinone. The major fatty acids were iso-C15 : 0 and anteiso-C15 : 0. The DNA G+C content was 37.9 mol%. A phylogenetic analysis based on 16S rRNA gene sequences showed that strain ISL-4T belonged to the genus Salegentibacter, exhibiting sequence similarity values of 95.9–98.6 % with respect to the type strains of recognized Salegentibacter species (with the exception of Salegentibacter catena). Low DNA–DNA relatedness values and differential phenotypic properties, together with its phylogenetic distinctiveness, demonstrated that strain ISL-4T represents a novel species of the genus Salegentibacter, for which the name Salegentibacter salinarum sp. nov. is proposed. The type strain is ISL-4T (=KCTC 12975T =CCUG 54354T).
-
-
-
Algoriphagus hitonicola sp. nov., isolated from an athalassohaline lagoon
More LessA Gram-negative, heterotrophic, aerobic, reddish-orange-pigmented, non-spore-forming, non-motile bacterial strain, designated 7-UAHT, was isolated from salty water from the athalassohaline lagoon at El Hito, located in central Spain. The strain grew optimally at 37 °C and pH 7.0 and in the presence of 2.5 % NaCl. A polyphasic taxonomic study was carried out in order to characterize the strain in detail. Phylogenetic analyses based on 16S rRNA gene sequence comparisons indicated that strain 7-UAHT clustered within the branch constituted by species of the genus Hongiella, which were recently transferred to the genus Algoriphagus. Analysis of the polar lipid profile and DNA G+C content also supported placement of strain 7-UAHT within the genus Algoriphagus. On the basis of its phenotypic, chemotaxonomic and phylogenetic distinctiveness, strain 7-UAHT represents a novel species of the genus Algoriphagus, for which the name Algoriphagus hitonicola sp. nov. is proposed. The type strain is 7-UAHT (=CECT 7267T =CCM 7449T).
-
-
-
Reclassification of Salegentibacter catena Ying et al. 2007 as Salinimicrobium catena gen. nov., comb. nov. and description of Salinimicrobium xinjiangense sp. nov., a halophilic bacterium isolated from Xinjiang province in China
More LessA Gram-negative, non-motile and moderately halophilic rod-shaped bacterium, designated strain BH206T, was isolated from a saline lake of Xinjiang province in China. The isolate showed catalase-positive and oxidase-negative reactions and did not reduce nitrate. Phylogenetic analysis based on 16S rRNA gene sequences showed that the isolate was most closely related to [Salegentibacter] catena HY1T with 95.8 % 16S rRNA gene sequence similarity, and formed a tight phyletic group with [Salegentibacter] catena HY1T with a bootstrap value of 99 % within the family Flavobacteriaceae. However, strain BH206T and [Salegentibacter] catena HY1T formed a phyletic lineage distinct from other Salegentibacter species. The 16S rRNA gene sequence similarities of strain BH206T with other related type species were lower than 94.6 %. On the basis of physiological and molecular properties, it is clear that [Salegentibacter] catena should be reclassified in the new genus Salinimicrobium as Salinimicrobium catena gen. nov., comb. nov. (type strain HY1T=CGMCC 1.6101T=JCM 14015T) and that strain BH206T represents a novel species within the genus Salinimicrobium, for which the name Salinimicrobium xinjiangense sp. nov. is proposed. The type strain of Salinimicrobium xinjiangense is BH206T (=KCTC 12883T=DSM 19287T).
-
-
-
Niabella soli sp. nov., isolated from soil from Jeju Island, Korea
More LessA dark yellow-coloured bacterium, JS13-8T, was isolated from a soil sample from Jeju Island, Republic of Korea. The cells were aerobic, Gram-negative, non-motile, short rods (0.5–0.7×0.8–1.4 μm). Growth occurred at 15–35 °C (optimally at 30 °C), at pH 5.0–8.0 (optimally at pH 6.0–7.0) and at 0–1 % NaCl (w/v). Flexirubin pigment was produced. A phylogenetic analysis based on 16S rRNA gene sequences revealed that strain JS13-8T was closely related to Niabella aurantiaca KACC 11698T (95.0 % sequence similarity). The major respiratory quinone system was MK-7 and the predominant cellular fatty acids were iso-C15 : 0, iso-C15 : 1 G, iso-C17 : 0 3-OH and summed feature 3. The DNA G+C content was 45 mol%. On the basis of the phylogenetic, physiological and chemotaxonomic data, strain JS13-8T represents a novel species of the genus Niabella, for which the name Niabella soli sp. nov. is proposed. The type strain is strain JS13-8T (=KACC 12604T=DSM 19437T).
-
-
-
Chryseobacterium soli sp. nov. and Chryseobacterium jejuense sp. nov., isolated from soil samples from Jeju, Korea
More LessTwo yellow-pigmented bacterial strains, JS6-6T and JS17-8T, isolated from soil samples from Jeju, Republic of Korea, were studied to determine their taxonomic positions. The cells of the two bacteria were aerobic, Gram-negative, non-motile, straight rods. 16S rRNA gene sequence analysis indicated that both isolates should be placed in the genus Chryseobacterium of the family Flavobacteriaceae. Their 16S rRNA gene sequences showed similarities of 93.7–97.5 % to those of type strains of the genus Chryseobacterium. The values for DNA–DNA relatedness between both strains and type strains of closely related Chryseobacterium species were below 34 %. The fatty acids of the novel strains were similar to those of species of the genus Chryseobacterium. Both strains had MK-6 as the predominant respiratory quinone. The DNA G+C contents of strains JS6-6T and JS17-8T were 39.9 and 41.4 mol%, respectively. Phylogenetic evidence, together with the DNA–DNA relatedness values and phenotypic characteristics, indicated that strains JS6-6T and JS17-8T represent two novel species of the genus Chryseobacterium, for which the names Chryseobacterium soli sp. nov. and Chryseobacterium jejuense sp. nov., respectively, are proposed. The type strains of Chryseobacterium soli sp. nov. and Chryseobacterium jejuense sp. nov. are JS6-6T (=KACC 12502T=DSM 19298T) and JS17-8T (=KACC 12501T=DSM 19299T), respectively.
-
-
-
Rudanella lutea gen. nov., sp. nov., isolated from an air sample in Korea
More LessAn orange-coloured bacterial strain, designated 5715S-11T, was isolated from an air sample in Suwon, Republic of Korea. The strain was strictly aerobic, Gram-negative, non-spore-forming, non-flagellated and rod-shaped, frequently forming filaments. Growth of the strain was observed at 4–40 °C (optimum, 30 °C), pH 6.0–8.0 (optimum, pH 7.0) and 0–2 % (w/v) NaCl. Phylogenetically, strain 5715S-11T was shown to be a member of the family Spirosomaceae within the phylum Bacteroidetes. Its closest relatives were Spirosoma linguale LMG 10896T (87.9 % 16S rRNA gene sequence similarity) and Larkinella insperata LMG 22510T (86.2 % 16S rRNA gene sequence similarity). Predominant cellular fatty acids were summed feature 3 (C16 : 1 ω7c and/or iso-C15 : 0 2-OH), C16 : 1 ω5c and iso-C15 : 0. Major polar lipids were phosphatidylethanolamine and unknown aminolipids and polar lipids. On the basis of the evidence from this polyphasic study, strain 5715S-11T is considered to represent a novel species of a new genus, for which the name Rudanella lutea gen. nov., sp. nov. is proposed. The type strain of Rudanella lutea is 5715S-11T (=KACC 12603T =DSM 19387T).
-
- Other Bacteria
-
-
Venenivibrio stagnispumantis gen. nov., sp. nov., a thermophilic hydrogen-oxidizing bacterium isolated from Champagne Pool, Waiotapu, New Zealand
More LessA novel thermophilic, hydrogen-oxidizing bacterium, designated strain CP.B2T, was isolated from a terrestrial hot spring in Waiotapu, New Zealand. Cells were motile, slightly rod-shaped, non-spore-forming and Gram-negative. Isolate CP.B2T was an obligate chemolithotroph, growing by utilizing H2 as electron donor and O2 as corresponding electron acceptor. Elemental sulfur (S0) or thiosulfate (
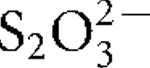 ) was essential for growth. Microbial growth occurred under microaerophilic conditions in 1.0–10.0 % (v/v) O2 [optimum 4–8 % (v/v) O2], between 45 and 75 °C (optimum 70 °C) and at pH values of 4.8–5.8 (optimum pH 5.4). The DNA G+C content was 29.3 mol%. 16S rRNA gene sequence analysis demonstrated that strain CP.B2T belonged to the order Aquificales, with a close phylogenetic relationship to Sulfurihydrogenibium azorense (94 % sequence similarity to the type strain). However, genotypic and metabolic characteristics differentiated the novel isolate from previously described genera of the Aquificales. Therefore, CP.B2T represents a novel species in a new genus, for which the name Venenivibrio stagnispumantis gen. nov., sp. nov. is proposed. The type strain of Venenivibrio stagnispumantis is CP.B2T (=JCM 14244T =DSM 18763T).
) was essential for growth. Microbial growth occurred under microaerophilic conditions in 1.0–10.0 % (v/v) O2 [optimum 4–8 % (v/v) O2], between 45 and 75 °C (optimum 70 °C) and at pH values of 4.8–5.8 (optimum pH 5.4). The DNA G+C content was 29.3 mol%. 16S rRNA gene sequence analysis demonstrated that strain CP.B2T belonged to the order Aquificales, with a close phylogenetic relationship to Sulfurihydrogenibium azorense (94 % sequence similarity to the type strain). However, genotypic and metabolic characteristics differentiated the novel isolate from previously described genera of the Aquificales. Therefore, CP.B2T represents a novel species in a new genus, for which the name Venenivibrio stagnispumantis gen. nov., sp. nov. is proposed. The type strain of Venenivibrio stagnispumantis is CP.B2T (=JCM 14244T =DSM 18763T).
-
-
-
‘Candidatus Phytoplasma omanense’, associated with witches'-broom of Cassia italica (Mill.) Spreng. in Oman
More LessSamples from plants of Cassia italica exhibiting typical witches'-broom symptoms (Cassia witches'-broom; CWB) were examined for the presence of plant pathogenic phytoplasmas by PCR amplification using universal phytoplasma primers. All affected plants yielded positive results. RFLP analyses of rRNA gene products indicated that the phytoplasmas detected were different from those described previously. Phylogenetic analysis of 16S rRNA gene sequences confirmed that CWB represents a distinct lineage and shares a common ancestor with ‘Candidatus Phytoplasma phoenicium’. Molecular comparison revealed that the 16S rRNA gene sequences of the four CWB strains (IM-1, IM-2, IM-3 and IM-4) identified in symptomatic C. italica samples were nearly identical (99.6–100 % similarity). The closest relatives were members of the pigeon pea witches'-broom phytoplasma ribosomal group (16SrIX; 95–97 % sequence similarity). On the basis of unique 16S rRNA gene sequences and biological properties, the phytoplasma associated with witches'-broom of C. italica in Oman represents a coherent but discrete novel phytoplasma, ‘Candidatus Phytoplasma omanense’, with GenBank/DDBJ/EMBL accession number EF666051 representing the reference strain.
-
- Proteobacteria
-
-
Hahella antarctica sp. nov., isolated from Antarctic seawater
More LessA Gram-negative, psychrotolerant, chemoheterotrophic, aerobic, cream-coloured bacterium, designated IMCC3113T, was isolated from coastal seawater from the Antarctic. On the basis of 16S rRNA gene sequence similarity analyses, the strain was most closely related to the type strains of Hahella chejuensis (93.0 %) and Hahella ganghwensis (92.1 %) in the Gammaproteobacteria. Phylogenetic investigations using 16S rRNA gene sequences showed that this Antarctic marine isolate formed a robust monophyletic clade with the two Hahella species but constituted a distinct phyletic line in the clade. The DNA G+C content of strain IMCC3113T was 56.4 mol% and the major respiratory quinone was Q-9. Several phenotypic and physiological characteristics, including the temperature range and NaCl optimum for growth, several enzyme activities and the cellular fatty acid composition, served to differentiate the strain from the two Hahella species. Therefore strain IMCC3113T represents a novel species of the genus Hahella, for which the name Hahella antarctica sp. nov. is proposed. The type strain is IMCC3113T (=KCCM 42675T =NBRC 102683T).
-
-
-
Helicobacter baculiformis sp. nov., isolated from feline stomach mucosa
More LessA Gram-negative, microaerophilic slender rod, measuring approximately 10 μm long and approximately 1 μm wide, isolated from the gastric mucosa of a cat and designated strain M50T, was subjected to a polyphasic taxonomic study. Despite its apparent lack of helical coils, the organism showed a corkscrew-like motion by means of multiple sheathed flagella located at both ends of the cell and by a periplasmic fibril coiled around the body. Strain M50T grew preferably on biphasic culture plates or on very moist agar. Coccoid forms predominated in cultures older than 4 days as well as in growth obtained on dry agar plates. The strain grew at 37 °C, but not at 25 or 42 °C and exhibited urease, oxidase and catalase activities. On the basis of 16S rRNA gene sequence analysis, the novel isolate was identified as a member of the genus Helicobacter and showed about 98 to 99 % sequence similarity to Helicobacter felis, Helicobacter bizzozeronii, Helicobacter salomonis, Helicobacter cynogastricus and ‘Candidatus Helicobacter heilmannii’, five highly related species previously detected in the feline or canine gastric mucosa. Protein profiling of strain M50T using SDS-PAGE revealed a pattern different from those of other Helicobacter species of mammalian gastric origin. Additionally, the urease and HSP60 gene sequences of strain M50T were different from those of H. felis, H. bizzozeronii, H. salomonis, H. cynogastricus and ‘Ca. H. heilmannii’. It is thus proposed that strain M50T (=LMG 23839T=CCUG 53816T) represents a novel species within this genus, for which the name Helicobacter baculiformis sp. nov. is proposed.
-
-
-
Lysobacter spongiicola sp. nov., isolated from a deep-sea sponge
More LessAn aerobic, Gram-negative bacterium, strain KMM 329T, was isolated from a deep-sea sponge specimen from the Philippine Sea and subjected to a polyphasic taxonomic investigation. Comparative 16S rRNA gene sequence analysis showed that strain KMM 329T clustered with the species of the genus Lysobacter. The highest level of 16S rRNA gene sequence similarity (97.0 %) was found with respect to Lysobacter concretionis KCTC 12205T; lower values (96.4–95.2 %) were obtained with respect to the other recognized Lysobacter species. The value for DNA–DNA relatedness between strain KMM 329T and L. concretionis KCTC 12205T was 47 %. Branched fatty acids 16 : 0 iso, 15 : 0 iso, 11 : 0 iso 3-OH and 17 : 1 iso were found to be predominant. Strain KMM 329T had a DNA G+C content of 69.0 mol%. On the basis of the phenotypic, chemotaxonomic, DNA–DNA hybridization and phylogenetic data, strain KMM 329T represents a novel species of the genus Lysobacter, for which the name Lysobacter spongiicola sp. nov. is proposed. The type strain is KMM 329T (=NRIC 0728T =JCM 14760T).
-
-
-
Brucella microti sp. nov., isolated from the common vole Microtus arvalis
More LessHolger C. Scholz, Zdenek Hubalek, Ivo Sedláček, Gilles Vergnaud, Herbert Tomaso, Sascha Al Dahouk, Falk Melzer, Peter Kämpfer, Heinrich Neubauer, Axel Cloeckaert, Marianne Maquart, Michel S. Zygmunt, Adrian M. Whatmore, Enevold Falsen, Peter Bahn, Cornelia Göllner, Martin Pfeffer, Birgit Huber, Hans-Jürgen Busse and Karsten NöcklerTwo Gram-negative, non-motile, non-spore-forming, coccoid bacteria (strains CCM 4915T and CCM 4916), isolated from clinical specimens of the common vole Microtus arvalis during an epizootic in the Czech Republic in 2001, were subjected to a polyphasic taxonomic study. On the basis of 16S rRNA (rrs) and recA gene sequence similarities, both isolates were allocated to the genus Brucella. Affiliation to Brucella was confirmed by DNA–DNA hybridization studies. Both strains reacted equally with Brucella M-monospecific antiserum and were lysed by the bacteriophages Tb, Wb, F1 and F25. Biochemical profiling revealed a high degree of enzyme activity and metabolic capabilities not observed in other Brucella species. The omp2a and omp2b genes of isolates CCM 4915T and CCM 4916 were indistinguishable. Whereas omp2a was identical to omp2a of brucellae from certain pinniped marine mammals, omp2b clustered with omp2b of terrestrial brucellae. Analysis of the bp26 gene downstream region identified strains CCM 4915T and CCM 4916 as Brucella of terrestrial origin. Both strains harboured five to six copies of the insertion element IS711, displaying a unique banding pattern as determined by Southern blotting. In comparative multilocus VNTR (variable-number tandem-repeat) analysis (MLVA) with 296 different genotypes, the two isolates grouped together, but formed a separate cluster within the genus Brucella. Multilocus sequence typing (MLST) analysis using nine different loci also placed the two isolates separately from other brucellae. In the IS711-based AMOS PCR, a 1900 bp fragment was generated with the Brucella ovis-specific primers, revealing that the insertion element had integrated between a putative membrane protein and cboL, encoding a methyltransferase, an integration site not observed in other brucellae. Isolates CCM 4915T and CCM 4916 could be clearly distinguished from all known Brucella species and their biovars by means of both their phenotypic and molecular properties, and therefore represent a novel species within the genus Brucella, for which the name Brucella microti sp. nov. with the type strain CCM 4915T (=BCCN 07-01T=CAPM 6434T) is proposed.
-
-
-
Paracoccus marinus sp. nov., an adonixanthin diglucoside-producing bacterium isolated from coastal seawater in Tokyo Bay
More LessTwo novel marine, Gram-negative, non-motile, catalase- and oxidase-positive, aerobic bacteria were isolated from coastal seawater in Tokyo Bay. Analysis of almost-complete 16S rRNA gene sequences showed that the two isolates are members of the genus Paracoccus, sharing highest 16S rRNA gene sequence similarity (96.5 %) with Paracoccus aminophilus NBRC 16710T. The DNA–DNA reassociation values between P. aminophilus NBRC 16710T and these isolates were only 10–20 %, in contrast to the high DNA relatedness between the two isolates (89 %). At least 1 % (w/v) NaCl was required for growth. Cellular fatty acid profiles revealed C18 : 1 ω7c as the major component and C10 : 0 3-OH as the major hydroxy fatty acid. Ubiquinone-10 was detected as the major respiratory quinone. The G+C content of the genomic DNA of both strains was 69 mol%. On the basis of DNA–DNA hybridization data and physiological and chemotaxonomic characteristics, it is proposed that these strains should be placed in a novel species, Paracoccus marinus sp. nov. The type strain is KKL-A5T (=NBRC 100637T =CIP 108500T); KKL-B9 (=NBRC 100640) is a reference strain.
-
-
-
Lysobacter capsici sp. nov., with antimicrobial activity, isolated from the rhizosphere of pepper, and emended description of the genus Lysobacter
More LessThe taxonomic position of a novel bacterial strain, YC5194T, with antimicrobial activity, isolated from the rhizosphere of pepper in Jinju, South Korea, was studied using a polyphasic approach. Cells of the strain were Gram-negative, rod-shaped, facultative anaerobes. It grew at a temperature of 15–37 °C (optimum 28 °C). Growth of the strain occurred between pH 5.5 and 8.5, with an optimum of pH 7.0–7.5. The strain inhibited mycelial growth of Pythium ultimum, Colletotrichum gloeosporioides, Fusarium oxysporum, Botrytis cinerea, Rhizoctonia solani and Botryosphaeria dothidea and growth of Bacillus subtilis. The G+C content of the total DNA was 65.4 mol%. The 16S rRNA gene sequence of the strain was most closely related to species of the genus Lysobacter (<94.0 to >99.0 % sequence similarity). Chemotaxonomic data (major quinone, Q-8; major polar lipids, diphosphatidylglycerol, phosphatidylethanolamine, phosphatidylglycerol and phosphatidyl-N-methylethanolamine; major fatty acids, iso-C15 : 0, summed feature 3, C16 : 0, iso-C17 : 1 ω9c and C18 : 1 ω7c) supported the affiliation of strain YC5194T to the genus Lysobacter. Phylogenetic analysis based on 16S rRNA gene sequences, DNA–DNA hybridization data and biochemical and physiological characteristics strongly supported the genotypic and phenotypic differentiation of strain YC5194T from species of Lysobacter with validly published names. Strain YC5194T therefore represents a novel species, for which the name Lysobacter capsici sp. nov. is proposed. The type strain is YC5194T (=KCTC 22007T =DSM 19286T).
-
-
-
Hydrogenophaga bisanensis sp. nov., isolated from wastewater of a textile dye works
More LessA Gram-negative, non-spore-forming, rod-shaped, Hydrogenophaga-like bacterial strain, K102T, was isolated from wastewater collected from a textile dye works in Korea and subjected to a polyphasic taxonomic study. Strain K102T grew optimally at pH 7.0–8.0 and 30–37 °C in the presence of 0.5 % (w/v) NaCl. It contained Q-8 as the predominant ubiquinone and C16 : 0, C16 : 1 ω7c and/or iso-C15 : 0 2-OH and C18 : 1 ω7c as the major fatty acids. The DNA G+C content was 64.8 mol%. A neighbour-joining phylogenetic tree based on 16S rRNA gene sequences showed that strain K102T fell within the radiation of the cluster comprising species of the genus Hydrogenophaga. Strain K102T exhibited 16S rRNA gene sequence similarity values of 95.9–98.9 % to the type strains of recognized Hydrogenophaga species. Levels of DNA–DNA relatedness between strain K102T and the type strains of its four phylogenetically most closely related species, together with differential phenotypic properties, revealed that strain K102T could be distinguished from all recognized species of the genus Hydrogenophaga. On the basis of phenotypic, phylogenetic and genetic data, strain K102T is considered to represent a novel species of the genus Hydrogenophaga, for which the name Hydrogenophaga bisanensis sp. nov. is proposed. The type strain is K102T (=KCTC 12980T =CCUG 54518T).
-
-
-
Allochromatium renukae sp. nov.
More LessAn ovoid to rod-shaped, phototrophic, purple sulfur bacterium, designated strain JA136T, was isolated in pure culture from brackish water near Kakinada, India, in a medium that contained 2 % NaCl (w/v). Cells were Gram-negative and motile by means of a single polar flagellum. Strain JA136T had no absolute salt requirement for growth but was able to tolerate up to 4 % NaCl (w/v). Intracellular photosynthetic membranes were of the vesicular type. Bacteriochlorophyll a and the carotenoid lycopene of the rhodopinal series were present as photosynthetic pigments. Strain JA136T was able to grow photolithoautotrophically and photolithoheterotrophically. There was no vitamin requirement for growth of strain JA136T. Phylogenetic analysis on the basis of 16S rRNA gene sequences indicated that strain JA136T clustered with species of the genus Allochromatium in the class Gammaproteobacteria. Strain JA136T showed highest 16S rRNA gene sequence similarity with the type strains of Allochromatium vinosum (97.0 %), Allochromatium minutissimum (95.8 %) and Allochromatium warmingii (90.0 %). Based on 16S rRNA gene sequence similarity data and morphological and physiological characteristics, strain JA136T was sufficiently distinct from recognized Allochromatium species to be described as representing a novel species of the genus, for which the name Allochromatium renukae sp. nov. is proposed. The type strain is JA136T (=JCM 14262T =DSM 18713T).
-
-
-
Collimonas arenae sp. nov. and Collimonas pratensis sp. nov., isolated from (semi-)natural grassland soils
More LessA polyphasic taxonomic study was performed to compare 26 novel bacterial isolates obtained from (semi-)natural grassland soils and a heathland soil in the Netherlands with 16 strains that had previously been assigned to the genus Collimonas. Genomic fingerprinting (BOX-PCR), whole-cell protein electrophoresis, matrix-assisted laser desorption ionization time-of-flight mass spectrometry of intact cells and physiological characterization (Biolog) of the isolates confirmed the existence of different strain clusters (A–D) within the genus Collimonas. Until now, only cluster C strains have been formally classified, as Collimonas fungivorans. In this study, DNA–DNA hybridizations were performed with a selection of strains representing the four clusters. The results showed that cluster B strains also belong to C. fungivorans and that strains of clusters A and D represent two novel species within the genus Collimonas. The latter novel species could be differentiated by means of phenotypic and genotypic characteristics and are classified as Collimonas arenae sp. nov. (cluster A; type strain Ter10T =LMG 23964T =CCUG 54727T) and Collimonas pratensis sp. nov. (cluster D; type strain Ter91T =LMG 23965T =CCUG 54728T).
-
-
-
Burkholderia sartisoli sp. nov., isolated from a polycyclic aromatic hydrocarbon-contaminated soil
More LessA Gram-negative, rod-shaped, aerobic bacterium, designated strain RP007T, was isolated from a polycyclic aromatic hydrocarbon-contaminated soil in New Zealand. Two additional strains were recovered from a compost heap in Belgium (LMG 18808) and from the rhizosphere of maize in the Netherlands (LMG 24204). The three strains had virtually identical 16S rRNA gene sequences and whole-cell protein profiles, and they were identified as members of the genus Burkholderia, with Burkholderia phenazinium as their closest relative. Strain RP007T had a DNA G+C content of 63.5 mol% and could be distinguished from B. phenazinium based on a range of biochemical characteristics. Strain RP007T showed levels of DNA–DNA relatedness towards the type strain of B. phenazinium and those of other recognized Burkholderia species of less than 30 %. The results of 16S rRNA gene sequence analysis, DNA–DNA hybridization experiments and physiological and biochemical tests allowed the differentiation of strain RP007T from all recognized species of the genus Burkholderia. Strains RP007T, LMG 18808 and LMG 24204 are therefore considered to represent a single novel species of the genus Burkholderia, for which the name Burkholderia sartisoli sp. nov. is proposed. The type strain is RP007T (=LMG 24000T =CCUG 53604T =ICMP 13529T).
-
-
-
Thalassobaculum litoreum gen. nov., sp. nov., a member of the family Rhodospirillaceae isolated from coastal seawater
More LessA Gram-negative, facultatively anaerobic strain with slightly curved and straight rod-shaped cells, strain CL-GR58T, was isolated from coastal seawater (near Gori, Korea). Analyses of the 16S rRNA gene sequence revealed that strain CL-GR58T belonged to the family Rhodospirillaceae with Azospirillum lipoferum as its closest relative (gene sequence similarity of 90.9 %). Phylogenetic analyses of the 16S rRNA gene sequences showed that strain CL-GR58T was not associated with any known genera in the family Rhodospirillaceae. The novel strain grew in the presence of 1–10 % sea salts, optimally at 30–35 °C and pH 8. The major cellular fatty acids consisted of C18 : 1 ω7c (48.5 %), C16 : 0 (14.8 %), C17 : 0 (12.2 %), C19 : 0 cyclo ω8c (6.3 %) and summed feature 3 (C16 : 1 ω7c and/or iso-C15 : 0 2-OH, 6.0 %). Among the phylogenetically related genera, the fatty acid C17 : 0 was found only in strain CL-GR58T. The DNA G+C content of the novel strain was 68.0 mol%. According to phylogenetic analyses of the 16S rRNA gene sequence, fatty acid content and the physiological data, strain CL-GR58T represents a novel species in a new genus of the family Rhodospirillaceae, for which the name Thalassobaculum litoreum gen. nov., sp. nov. is proposed. The type strain of the type species is CL-GR58T (=KCCM 42674T=DSM 18839T).
-
-
-
Pseudacidovorax intermedius gen. nov., sp. nov., a novel nitrogen-fixing betaproteobacterium isolated from soil
More LessA Gram-negative, short rod-shaped, nitrogen-fixing bacterium (CC-21T) was isolated from a soil sample collected from the regional agricultural research station in Kaohsiung County (Taiwan). Using 16S rRNA gene sequence analysis, it could be clearly demonstrated that this isolate was novel: it showed <97 % similarity to species of the genera Acidovorax, Alicycliphilus, Giesbergeria, Simplicispira and Diaphorobacter. The organism used several organic acids, but only a few sugars as substrates. The fatty acid profile differed from those reported for members of the genera Acidovorax, Alicycliphilus, Giesbergeria, Simplicispira and Diaphorobacter. On the basis of 16S rRNA gene sequence analysis in combination with physiological data, strain CC-21T represents a novel species in a new genus, for which the name Pseudacidovorax intermedius gen. nov., sp. nov. is proposed; the type strain is CC-21T (=CCUG 54492T=CIP 109510T).
-
-
-
Jannaschia pohangensis sp. nov., isolated from seashore sand in Korea
More LessA Gram-negative, aerobic bacterium, H1-M8T, was isolated from seashore sand in Korea and then characterized using a polyphasic approach. Cells were short rods (0.7–1.0×1.5–2.0 μm) and were motile (each cell having at least one flagellum). Colonies were light-brown, non-pigmented, circular and convex with clear margins. Growth of the strain was observed at 5–35 °C, pH 6.0–9.0 and NaCl concentrations up to 8.4 % (w/v). Phylogenetic analysis of the 16S rRNA gene sequence revealed a clear affiliation between the novel strain and members of the genus Jannaschia. The sequence similarities between H1-M8T and type strains of the genus Jannaschia ranged from 97.0 to 97.8 %. However, DNA–DNA hybridizations between the isolate and type strains of other related species produced low values (21–38 %). The major isoprenoid quinone was Q-10 and the predominant cellular fatty acids were 18 : 1ω7c (68.2 %) and 18 : 0 (10.5 %). The G+C content of the DNA was 63.6 mol%. On the basis of physiological, biochemical and chemotaxonomic traits and data from the comparative 16S rRNA sequence analysis, strain H1-M8T represents a novel species of the genus Jannaschia, for which the name Jannaschia pohangensis sp. nov. is proposed. The type strain is H1-M8T (=KACC 11609T =DSM 19073T).
-
- Eukaryotic Micro-Organisms
-
-
Saccharomyces arboricolus sp. nov., a yeast species from tree bark
More LessThree ascomycetous yeast strains, H-6T, ZX-15 and ZX-20, isolated from the bark of two tree species of the family Fagaceae collected from different regions of China, formed unconjugated and persistent asci containing two to four globose ascospores. 26S rDNA D1/D2 domain and internal transcribed spacer (ITS) region (including 5.8S rDNA) sequence analysis showed that they were closely related to the currently accepted Saccharomyces species with strong support. Comparisons of the rDNA sequences, electrophoretic karyotypes and physiological characters indicated that the three strains represent a novel species in the genus Saccharomyces. The name Saccharomyces arboricolus sp. nov. was proposed for the novel species, with H-6T (=AS 2.3317T=CBS 10644T) isolated from the bark of Quercus fabri as the type strain.
-
-
-
Candida phangngensis sp. nov., an anamorphic yeast species in the Yarrowia clade, isolated from water in mangrove forests in Phang-Nga Province, Thailand
More LessTwo yeast strains (TM2-16 and PT1-17T) were isolated by membrane filtration from samples of estuarine water collected from two mangrove forests, in Khao Lumpee-Haad Thaimueang National Park and Mu Ko Ra-Ko Prathong National Park, Phang-Nga Province, Thailand. Analysis of the D1/D2 domain of the large-subunit rDNA sequences revealed that the sequences of the two strains were identical. The closest species in terms of pairwise sequence similarity was Candida galli, but the level of nucleotide substitutions (13.2 %) was sufficient to justify the description of a separate species. Phylogenetic analysis demonstrated that the two strains occupy a basal position with respect to Yarrowia lipolytica and C. galli of the Yarrowia clade, supported by a high bootstrap value. The two strains showed identical phenotypic characteristics, including proliferation by multilateral budding, absence of ascospores and ballistoconidia and negative Diazonium blue B and urease reactions. The major ubiquinone was Q-9. On the basis of the above findings, these two strains were assigned to a single novel species of the genus Candida, for which the name Candida phangngensis sp. nov. is proposed. The type strain is PT1-17T (=BCC 21231T =NBRC 101967T =CBS 10407T).
-
-
-
Kurtzmaniella gen. nov. and description of the heterothallic, haplontic yeast species Kurtzmaniella cleridarum sp. nov., the teleomorph of Candida cleridarum
More LessThe teleomorph of Candida cleridarum was discovered through the detection of conjugation between isolates of a large collection from the nitidulid beetles of the genus Carpophilus found in the flowers of various cacti in Arizona, USA. The previous oversight of the sexual cycle of this yeast is attributed to the inequality (ca. 5 : 1) of the two mating types. Extensive conjugation between compatible mating types is observed after overnight incubation on 5 % malt agar, followed after 3–5 days by the formation of mature asci. The hat-shaped ascospores are reminiscent of those seen in Kodamaea species, which are members of the same guild. However, published analyses of D1/D2 large subunit rDNA sequences indicate an affinity with the genus Debaryomyces. As the latter is polyphyletic and morphologically heterogeneous, and in view of the distinct life cycle of the new teleomorph, the new genus Kurtzmaniella is described with a novel species, Kurtzmaniella cleridarum sp. nov. Given the close relatedness of Kurtzmaniella cleridarum sp. nov. to Candida quercitrusa, Candida oleophila and Candida railenensis, for which several natural isolates were available, strains of these species were mixed in pairs under the conditions found favourable for the former. Conjugation was not detected in those species. The type strain of Kurtzmaniella cleridarum sp. nov. is UWOPS 99-101.1T (=CBS 8793T=NRRL Y-48386T, h+ ), type of Candida cleridarum. The allotype is UWOPS 07-123.1 (=CBS 10688=NRRL Y-48387, h− ).
-
- Other Gram-Positive Bacteria
-
-
Paenibacillus forsythiae sp. nov., a nitrogen-fixing species isolated from rhizosphere soil of Forsythia mira
More LessA nitrogen-fixing bacterium, designated strain T98T, was isolated from rhizosphere soil of Forsythia mira. Phylogenetic analysis based on a fragment of the nifH gene and the full-length 16S rRNA gene sequence revealed that strain T98T was a member of the genus Paenibacillus. High levels of 16S rRNA gene similarity were found between strain T98T and Paenibacillus durus ATCC 35681T (97.0 %), Paenibacillus sabinae DSM 17841T (98.3 %) and Paenibacillus zanthoxyli DSM 18202T (96.8 %). Levels of 16S rRNA gene sequence similarity between strain T98T and the type strains of other recognized members of the genus Paenibacillus were below 97.0 %. Levels of DNA–DNA relatedness between strain T98T and P. durus ATCC 35681T, P. sabinae DSM 17841T and P. zanthoxyli DSM 18202T were 27.6, 30.0 and 32.1 %, respectively. The DNA G+C content of strain T98T was 50.4 mol%. The major fatty acids were anteiso-C15 : 0, C16 : 0 and iso-C16 : 0. On the basis of its phenotypic characteristics and levels of DNA–DNA hybridization, strain T98T is considered to represent a novel species of the genus Paenibacillus, for which the name Paenibacillus forsythiae sp. nov. is proposed. The type strain is T98T (=CCBAU 10203T =DSM 17842T).
-
-
-
Virgibacillus chiguensis sp. nov., a novel halophilic bacterium isolated from Chigu, a previously commercial saltern located in southern Taiwan
More LessA Gram-positive, motile, endospore-forming, irregular rod-shaped (0.7–0.9×2.5–5.0 μm), halophilic bacterial strain, NTU-101T, was isolated from Chigu saltern in southern Taiwan, previously used as a salt production field. The isolate was characterized taxonomically based on biochemical and molecular approaches. It grows optimally at 40 °C and in the presence of 5–10 % NaCl. Strain NTU-101T has cell-wall peptidoglycan based on meso-diaminopimelic acid and MK-7 as the predominant menaquinone. Major polar lipids are phosphatidylglycerol, diphosphatidylglycerol and phosphatidylethanolamine. Anteiso-C15 : 0, iso-C15 : 0 and anteiso-C17 : 0 are the major fatty acids. The DNA G+C content was 37.3 mol%. Phylogenetic analyses based on 16S rRNA gene sequences showed its affiliation to the genus Virgibacillus. DNA–DNA relatedness values between strain NTU-101T and Virgibacillus dokdonensis and Virgibacillus pantothenticus were 17.5 and 21.5 %, respectively. Based on the phenotypic and phylogenetic properties, strain NTU-101T (=BCRC 17637T=CGMCC 1.6496T) was classified as a novel strain of Virgibacillus species, for which the name Virgibacillus chiguensis sp. nov. is proposed.
-
-
-
Bacillus butanolivorans sp. nov., a species with industrial application for the remediation of n-butanol
More LessFour bacterial strains, designated K9T, K105, K1012A and K101, were isolated from soil in Lithuania. All these strains could use n-butanol as a sole carbon source. The strains grew in a medium containing 12–120 mM n-butanol. The strains were strictly aerobic, Gram-positive endospore-formers. The best growth was achieved at 25 °C and pH 7.0 in medium containing 1 % (w/v) NaCl. The strains showed identical profiles of 16S–23S rRNA internal transcribed spacer PCR and nearly identical 16S rRNA gene PCR-RFLP electrophoretic patterns and physiological characteristics, demonstrating their relationship at the species level. The cellular fatty acid profile of K9T consisted of significant amounts of the C15 branched-chain fatty acids iso-C15 : 0 (16.78 %) and anteiso-C15 : 0 (45.80 %). The diagnostic cell-wall diamino acid was meso-diaminopimelic acid. The 16S rRNA gene sequence of K9T showed the highest similarity to the sequences of Bacillus simplex DSM 1321T and Bacillus muralis LMG 20238T (98.3 and 97.7 %, respectively). The DNA G+C content was 37.4 mol%. Studies of DNA–DNA relatedness, morphological, physiological and chemotaxonomic analyses and phylogenetic data based on 16S rRNA gene sequencing allowed strains K9T, K105, K1012A and K101 to be described as members of a novel species of the genus Bacillus, for which the name Bacillus butanolivorans sp. nov. is proposed. The type strain is K9T (=DSM 18926T =LMG 23974T).
-
- Evolution, Phylogeny And Biodiversity
-
-
-
Genetic and morphological characterization of Rivularia and Calothrix (Nostocales, Cyanobacteria) from running water
More LessIn this study, a polyphasic approach was adopted to investigate natural freshwater (river and stream) samples of Rivularia colonies and isolated strains of cyanobacteria with a high degree of trichome tapering (genera Rivularia and Calothrix). Analysis of the phycocyanin (PC) operon and the intervening intergenic spacer (cpcBA-IGS) and 16S rRNA gene sequences were used for genetic characterization. In addition, a molecular fingerprinting method, temperature-gradient gel electrophoresis, which allows sequence-dependent separation of PCR products, was used to assess genotypic diversity in environmental samples and isolated strains. The results showed a high variability of the PC-IGS among the genotypes that was not associated with the morphologies observed. This study underlines the importance of choosing a low-nutrient-content culture medium, especially one with a low phosphorus concentration, for studying typical morphological features of Rivularia for taxonomic purposes. Molecular fingerprinting methods and morphological analyses confirmed the diversity in Rivularia colonial structure and trichome features corresponding to genetic diversity within a single colony. Phylogenetic analysis of cpcBA-IGS was largely consistent with that obtained from 16S rRNA gene sequence analysis and confirmed the high level of divergence between genotypes. The sequences of Rivularia and Calothrix from this study and database sequences showed great heterogeneity and were clearly not monophyletic. The results of this genetic and morphological study of field samples and fresh isolates indicated that the current classification of these genera needs to be revised.
-
-
- International Committee On Systematics Of Prokaryotes
-
- Taxonomic Note
-
-
Proposal of Yaniellaceae fam. nov., Yaniella gen. nov. and Sinobaca gen. nov. as replacements for the illegitimate prokaryotic names Yaniaceae Li et al. 2005, Yania Li et al. 2004, emend Li et al. 2005, and Sinococcus Li et al. 2006, respectively
More LessThe prokaryotic generic names Yania Li et al. 2004 and Sinococcus Li et al. 2006 are illegitimate because they are later homonyms of the names Yania Roewer 1919 (Opiliones, Arachnida, Arthropoda, Animalia), Yania Huang 1997 (Lepidoptera: Hesperiidae) and Sinococcus Wu and Zheng 2000 (Homoptera: Coccomorpha) [Principle 2 of the Bacteriological Code (1990 Revision)]. Therefore, new generic names, Yaniella gen. nov. and Sinobaca gen. nov., are proposed for these taxa. In addition, a new family name, Yaniellaceae fam. nov., is proposed to accommodate Yaniella gen. nov. As a result, new combinations are required for the species to replace the illegitimate species names.
-
Volumes and issues
-
Volume 74 (2024)
-
Volume 73 (2023)
-
Volume 72 (2022 - 2023)
-
Volume 71 (2020 - 2021)
-
Volume 70 (2020)
-
Volume 69 (2019)
-
Volume 68 (2018)
-
Volume 67 (2017)
-
Volume 66 (2016)
-
Volume 65 (2015)
-
Volume 64 (2014)
-
Volume 63 (2013)
-
Volume 62 (2012)
-
Volume 61 (2011)
-
Volume 60 (2010)
-
Volume 59 (2009)
-
Volume 58 (2008)
-
Volume 57 (2007)
-
Volume 56 (2006)
-
Volume 55 (2005)
-
Volume 54 (2004)
-
Volume 53 (2003)
-
Volume 52 (2002)
-
Volume 51 (2001)
-
Volume 50 (2000)
-
Volume 49 (1999)
-
Volume 48 (1998)
-
Volume 47 (1997)
-
Volume 46 (1996)
-
Volume 45 (1995)
-
Volume 44 (1994)
-
Volume 43 (1993)
-
Volume 42 (1992)
-
Volume 41 (1991)
-
Volume 40 (1990)
-
Volume 39 (1989)
-
Volume 38 (1988)
-
Volume 37 (1987)
-
Volume 36 (1986)
-
Volume 35 (1985)
-
Volume 34 (1984)
-
Volume 33 (1983)
-
Volume 32 (1982)
-
Volume 31 (1981)
-
Volume 30 (1980)
-
Volume 29 (1979)
-
Volume 28 (1978)
-
Volume 27 (1977)
-
Volume 26 (1976)
-
Volume 25 (1975)
-
Volume 24 (1974)
-
Volume 23 (1973)
-
Volume 22 (1972)
-
Volume 21 (1971)
-
Volume 20 (1970)
-
Volume 19 (1969)
-
Volume 18 (1968)
-
Volume 17 (1967)
-
Volume 16 (1966)
-
Volume 15 (1965)
-
Volume 14 (1964)
-
Volume 13 (1963)
-
Volume 12 (1962)
-
Volume 11 (1961)
-
Volume 10 (1960)
-
Volume 9 (1959)
-
Volume 8 (1958)
-
Volume 7 (1957)
-
Volume 6 (1956)
-
Volume 5 (1955)
-
Volume 4 (1954)
-
Volume 3 (1953)
-
Volume 2 (1952)
-
Volume 1 (1951)
Most Read This Month


